Results of this study provide new evidence of the influence\r\nof both abiotic and biotic factors on the gut bacteriome of aquatic species of\r\ncommercial interest, like Pacific white shrimp. Photo b...
Results of this study provide new evidence of the influence\r\nof both abiotic and biotic factors on the gut bacteriome of aquatic species of\r\ncommercial interest, like Pacific white shrimp. Photo by Darryl Jory.
Among the infectious diseases of penaeid shrimp, the white\r\nspot syndrome (WSS) and the acute hepatopancreatic necrosis disease (AHPND, previously\r\nreferred as “early mortality syndrome” or EMS) are currently the most serious\r\nthreats to shrimp farmers. WSS is caused by the white spot syndrome virus\r\n(WSSV), whereas the AHPND is an emerging bacteriosis caused by virulent strains\r\nof Vibrio parahaemolyticus and V. harveyi. The adoption of\r\non-farm biosecurity practices is necessary to limit the pathogen entrance into\r\nthe culture systems.
Among on-shrimp farm biosecurity practices, the biofloc super-intensive technology (BFT) is a promising alternative culture system. It is widely believed that BFT culture improves the crustacean immunity leading to\r\nhigh survival rates even under bacterial and viral infections. Although the\r\nmechanisms underlying shrimp robustness are not yet understood, a continuous immunostimulation\r\ncondition is rather expected considering the abundance of microbial-associated\r\nmolecular patterns (MAMPs) present in BFT systems which can activate innate\r\nimmune responses.
Since the aquatic environment can influence the microbiota\r\ncomposition and abundance, studies focusing the BFT contribution on the\r\nestablishment of shrimp intestinal microbiota are highly required. In addition,\r\nit is now well established that commensal microbiota is essential for the\r\ncorrect functionality of the host physiology. Surprisingly, the\r\ncharacterization of the microbial communities present in the digestive tract of\r\ncultured shrimp species has been only recently uncovered.
To our knowledge, only one report regarding the description\r\nof the intestinal bacterial communities of a penaeid species – the Pacific blue\r\nshrimp (Litopenaeus stylirostris) – reared in a BFT system is available in the\r\nliterature. In addition, nothing is known about the bacteriome (a specialized\r\norgan with specialized cells that provide shelter and nutrients to the bacteria\r\nwhile protecting the host animal) plasticity in shrimp infected by the WSSV,\r\none of the most important pathogens in shrimp farming.
This article – adapted and summarized from the original publication –\r\nwas designed to characterize the abundance and composition of the intestinal\r\nbacterial communities of the most important penaeid species, the Pacific white\r\nshrimp (L. vannamei) reared in BFT and a clear seawater system. Likewise, the\r\nplasticity of the midgut bacteriome from shrimp challenged by WSSV was\r\ninvestigated. Our results bring new evidence of the influence of the biofloc\r\nculture and the viral challenge on the shrimp bacteriome, providing new\r\ninsights into future studies regarding the role of microbiota on the intestinal\r\nimmunity of cultured penaeid.
Study setup
L. vannamei postlarvae\r\n(five days old, PL5) from a commercial shrimp hatchery (Aquatec LTDA\r\nCanguaratema, Rio Grande do Norte, Brazil) were used in this study at the\r\nLaboratory of Marine Shrimp (Federal University of Santa Catarina,\r\nFlorianópolis, Brazil). The biofloc culture was initially established in a\r\n50-cubic-meter matrix tank and the experimental design is shown in Fig. 1.
Shrimp PLs were randomly stocked into eight, 9-cubic-meter\r\nindoor tanks, with four BFT tanks and four clear seawater system tanks, at an\r\ninitial stocking density of 300 and 20 PL5 per cubic meter, respectively. The\r\ntanks were continuously aerated (dissolved oxygen > 5 mg/L) and kept under\r\ncontrolled temperature (29 ± 1 degrees-C) and salinity (34 to 35 ppt).
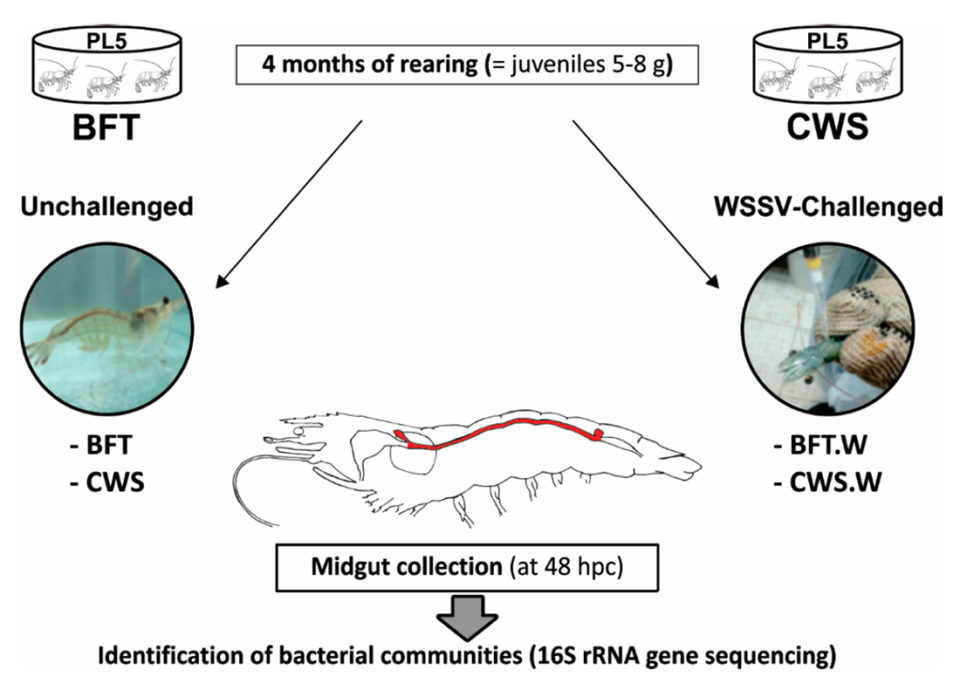
Fig. 1: Shrimp PL5 were cultured for four months in two\r\nculture systems, Biofloc Technology (BFT; 4 tanks) and clear seawater system\r\n(CWS; 4 tanks), at an initial stocking density of 300 and 20 animals per cubic\r\nmeter, respectively. Cultured, white spot syndrome virus (WSSV)-free juvenile\r\nshrimp (5 to 8 grams) from each treatment (n = 80) were then individually\r\nchallenged with WSSV via oral route (5 × 106 genome viral copies). At 48 hours\r\npost-challenge (hpc), the midguts from unchallenged (BFT and CWS) and\r\nWSSV-challenged (BFT.W and CWS.W) shrimp (n = 40/treatment) were collected and\r\nprocessed for 16S RNA gene sequencing.
Post-larvae were fed four times a day with a commercial diet\r\n(Guabi Potimar), and tank water was exchanged at 80 percent daily. After four\r\nmonths, when the shrimp had become 5- to 8-gram juveniles, around 30 percent of\r\nthe animals from each tank/group were randomly selected to confirm that the\r\nshrimp were free of WSSV by using the nested-PCR assay, and then 120 animals\r\nfrom each culture system were transferred to the Laboratory of Immunology\r\nApplied to Aquaculture (Federal University of Santa Catarina, Florianópolis,\r\nBrazil) for various tests and analyses.
For a detailed description of the experimental design; WSSV\r\nper os (oral) challenge and midgut collection; genomic DNA (gDNA) extraction;\r\n16S rRNA gene library preparation and high throughput sequencing; and sequence\r\ndata analysis, refer to the original publication.
Results and\r\ndiscussion
Our study explored the bacterial communities’ dynamics\r\n(abundance and phylogenetic composition) in the shrimp midgut in response to\r\ntwo important abiotic and biotic factors related to shrimp farming (culture\r\nsystem and viral infection) by assessing 16S rRNA gene sequencing [this technique\r\nis used in reconstructing phylogenies – evolutionary histories – due to the\r\nslow rates of evolution of this region of the gene]. We generated two bacterial\r\n16S rRNA gene libraries from the midguts of shrimp cultured in a BioFloc\r\nTechnology (library “BFT”) and in a clear seawater system (library “CWS”).\r\nLikewise, two other libraries were generated from the midguts of shrimp\r\nchallenged with the WSSV using a per os method (libraries “BFT.W” and “CWS.W”).
Analysis of Venn diagrams [which show all possible logical\r\nrelations between a finite collection of different sets] revealed significant\r\ndifferences in the frequency distribution of bacterial operational taxonomic\r\nunits (OTU; operational definition used to classify groups of closely\r\nrelated individuals) according to the culture system (BFT and CWS) and viral\r\nchallenge (Fig. 2). The midguts of animals reared in BFT exhibited a larger\r\nnumber of OTUs when compared with those from CWS. Five-hundred and seventy-one\r\nOTUs were exclusively found in shrimp reared in bioflocs (361 in BFT, 111 in\r\nBFT.W and 99 in both groups), whereas 298 OTUs were exclusive from shrimp\r\nreared in clear seawater (162 in CWS, 121 in CWS.W and 15 in both conditions).
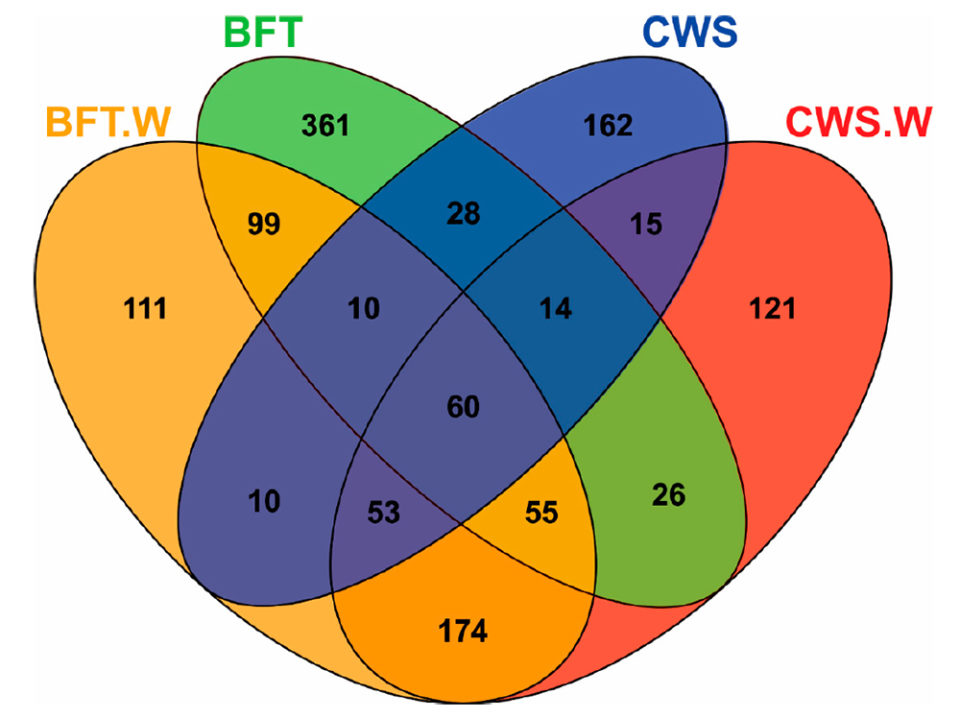
Fig. 2: Venn diagram showing unique and shared operational\r\ntaxonomic units (OTUs) of midgut bacteria of Litopenaeus vannamei reared\r\nin biofloc (BFT) and clear seawater (CWS) and challenged by the WSSV (BFT.W and\r\nCWS.W), by using a per os method.
The higher number of exclusive OTUs from the BFT samples\r\ncould reflect the diversity microbioma of the biofloc environment. The viral\r\nchallenge led to the appearance of exclusive OTUs in each rearing condition:\r\n111 OTUs for BFT.W and 121 for CWS.W. Furthermore, exclusive OTUs (n = 174)\r\nwere shared only by the challenged animals of both rearing conditions. These\r\nfindings suggest that this bacterial community displacement in the midgut is\r\nrelated to the virus presence. Finally, 60 OTUs were shared among all samples,\r\nwhich represented 4.61 percent of the total OTUs. This bacteria subset present\r\nin all groups could represent relevant microorganisms to the fundamental\r\nstructure and function of the shrimp intestinal microbiota. Additional studies\r\non microbiota shrimp-virus interaction related to the BFT rearing system\r\ndeserve to be investigated in the future.
Regarding the influence of rearing conditions on the\r\nbacterial communities of shrimp midgut, it is widely believed that the\r\ncommensal microbiota composition in adult arthropods [invertebrate animals with\r\nan exoskeleton (external skeleton), like insects and crustaceans] appears to be\r\nintimately related to the initial exposure to microorganisms during their early\r\nlife. We therefore cultured our shrimp in BFT and CWS for four months (PL5 to 5\r\nto 8 grams) before the lab testing. In our analysis, the resulting high-quality\r\n16S rRNA gene sequences were classified into 33 prokaryotic phyla that belong\r\nto the domain Bacteria. The most representative phyla identified in the L.\r\nvannamei midgut were Proteobacteria, Bacteroidetes, Actinobacteria and\r\nFirmicutes. However, the frequency distribution of the intestinal bacterial\r\ncommunities differed according to the rearing system (Fig. 3).
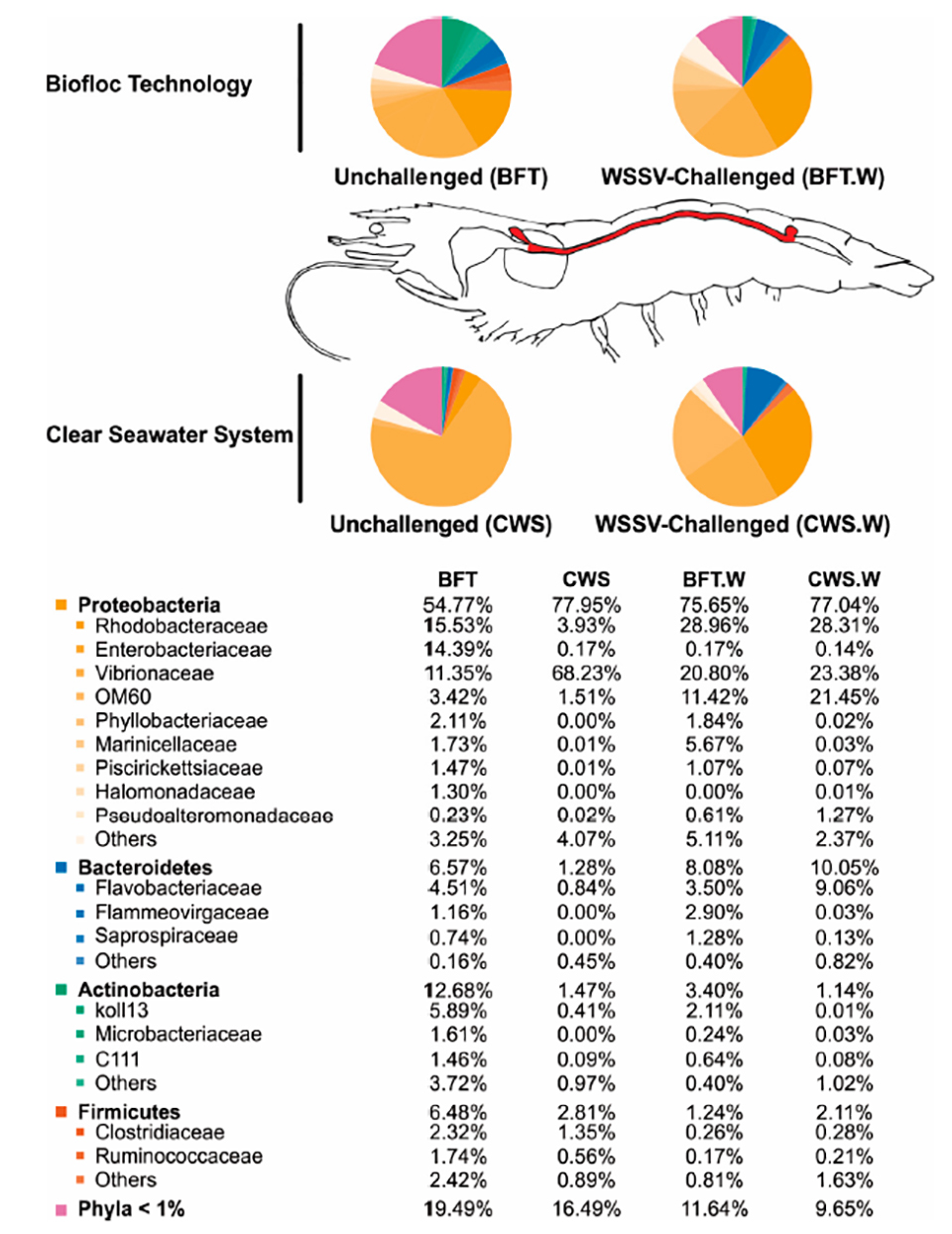
Fig. 3: Relative abundance of the most prevalent bacterial\r\nphyla and families identified in the midgut of L. vannamei (highlighted in\r\nred in the not-to-scale image) reared in biofloc (BFT) and clear seawater\r\n(CWS), and at 48 hours after an oral challenge with white spot syndrome virus (BFT.W\r\nand CWS.W).
The genus Vibrio is composed of fast-growing\r\naquatic bacteria able to colonize the digestive tract of different animals,\r\nincluding penaeids. Many Vibrio species are opportunistic pathogens\r\nin shrimp under stressful conditions, such as poor nutrition, low water\r\nquality, and immune depression. In cultured shrimp, this bacterial group has\r\nbeen repeatedly implicated in gastro-intestinal diseases, leading to high\r\nshrimp mortalities. Microorganisms present in BFT water could act against\r\npathogenic bacteria by competition for substrate and nutrients, producing\r\ninhibitory compounds, and interfering in the bacterial quorum-sensing [a system\r\nof stimuli and response correlated to population density, used by many bacteria\r\nto coordinate gene expression according to the density of their local\r\npopulation] communication.
Interestingly, two Proteobacteria families, Rhodobacteraceae\r\nand Enterobacteriaceae, were more abundant in the midgut of shrimp reared in\r\nBFT than in CWS, representing around 15 percent in BFT and less than 4 percent\r\nin CWS. The role of this bacterial family in the shrimp intestinal microbiota\r\nis not well understood, but it is believed that the BFT system can favor the\r\npresence of this bacteria family due to its high concentration of suspended solid\r\nthat can be used as growth sites by Rhodobacteraceae. Family members can\r\nestablish an antagonistic activity limiting the survival of pathogenic Vibrio.\r\nTherefore, we could hypothesize that the higher abundance of Rhodobacteraceae\r\nin the midgut of L. vannamei reared in BFT could be associated with a\r\nlower abundance of Vibrionaceae.
Our findings indicate that the bacteriome of shrimp reared\r\nin BFT was more diversified and richer when compared to that from animals\r\nreared in clear seawater, where the predominant bacterial community was\r\nVibrionaceae. The water from BFT is especially rich in organic matter and\r\nsuspended particles, which can favor bacteria that use organic matter and\r\nnitrogen compounds for growth. In addition, the BFT rearing system apparently causes\r\nimportant modifications in shrimp midgut microbiota compared to CWS, which\r\ncorroborates the fact that the microbiota from the digestive tract of the\r\naquatic animals is directly influenced by the environment. In addition, the\r\nidea of considering the BFT as a “natural probiotic system” has important\r\nconsequences to the intestinal microbiota. The BFT could act internally and/or\r\nexternally to the shrimp body, an effect promoted by large groups of\r\nmicroorganisms, but mainly bacteria.
Regarding the beneficial effects of the BFT system, bioflocs\r\ncan act as immunostimulants and enhancing the shrimp innate immune system, even\r\naltering the expression of genes related to the shrimp immune response, which\r\ncould be attributed to the ability of the BFT to induce changes in shrimp\r\nmicrobiota. Overall, our results show that the shrimp gut microbiota is altered\r\nby the rearing environment.
Regarding the shrimp intestinal microbiota plasticity in\r\nresponse to a viral challenge, the most impacting results of our study were to\r\ncharacterize the bacterial communities shift in the shrimp midgut challenged by\r\nWSSV. To the best of our knowledge, there is only one study on the shift of gut\r\nmicrobiota in response to the WSSV infection, evaluated in the Chinese mitten\r\ncrab. Ours is the first study investigating the effect of a viral pathogen on\r\nthe intestinal microbiota of a penaeid species reared in a BFT system.
Perspectives
By using a high-throughput sequencing technology, we have\r\ncharacterized the intestinal bacteriome of the most important cultivated shrimp\r\nspecies, L. vannamei, and assessed the influence of the BFT rearing and of\r\nthe WSSV challenge on the composition and abundance of the bacterial\r\ncommunities.
The bacterial composition from the midgut of shrimp reared\r\nin bioflocs was richer and more diverse than that from clear seawater. The\r\npredominant bacterial group belonged to the phylum Proteobacteria\r\n(Rhodobacteraceae, Enterobacteriaceae and Vibrionaceae), followed by the phyla\r\nBacteroidetes (Flavobacteriaceae), Actinobacteria and Firmicutes. Vibrionaceae\r\nwas more abundant in the CWS group than in BFT-reared shrimp (68.23 percent and\r\n11.35 percent from total bacterial communities, respectively).
The bacterial composition of L. vannamei midgut\r\nwas affected by the WSSV challenge. Vibrionaceae was the most affected\r\nbacterial family and its abundance doubled in the midgut of BFT-reared shrimp\r\nafter viral challenge, while in CWS-reared shrimp decreased drastically. In\r\naddition, the WSSV challenge apparently led to a more homogeneous distribution\r\nof bacterial population composition, as Rhodobacteraceae, Enterobactereceae and\r\nVibrionaceae, in the midgut of shrimp reared in both culture systems, BFT and\r\nCWS. The changes in the gut bacteria diversity associated to the WSSV challenge\r\ncould indicate a displacement in the intestinal microbial communities leading\r\nto the dysbiosis (a microbial imbalance or maladaptation on or inside the body,\r\nlike an impaired microbiota) condition.
Knowing the intestinal bacterial populations of shrimp\r\nreared in BFT and during WSSV infection is a relevant step to understanding the\r\nrole of intestinal bacteriome microbiota on crustacean immune defenses against\r\nviral diseases. Although the molecular mechanisms involved in the control and\r\nregulation of the shrimp gut microbiota is still largely unknown, the\r\nenvironmental conditions and the presence of infectious agents proved to be\r\ndecisive factors influencing both the diversity and abundance of the bacterial\r\ncommunities.
\r\n\r\n\r\n\r\n\r\n\r\n\r\n\r\n\r\n\r\n\r\n\r\n\r\n\r\n\r\n\r\n\r\n\r\n\r\n\r\n\r\n\r\n\r\n\r\n\r\n\r\n\r\n\r\n\r\n\r\n\r\n\r\n\r\n\r\n\r\n\r\n\r\n\r\n\r\n\r\n\r\n\r\n\r\n\r\n\r\n\r\n\r\n\r\n\r\n\r\n\r\n\r\n\r\n\r\n\r\n\r\n\r\n\r\n
With these 16S rRNA sequencing data in hand, and given that\r\npenaeid shrimp is an excellent model for functional genomic studies, we can now\r\ninvestigate shrimp in host-microbiota interactions, and also the role of the\r\ncommensal microbiota in the regulation of the shrimp gut immunity.
Source : Global Aquaculture Alliance

Ditulis oleh
Tim Minapoli
Kontributor
Pakar di bidang akuakultur dengan pengalaman lebih dari 15 tahun. Aktif berkontribusi dalam pengembangan industri perikanan Indonesia.
